|
|||||
|
|
|||||
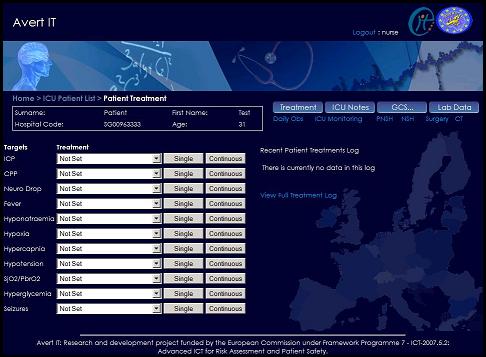
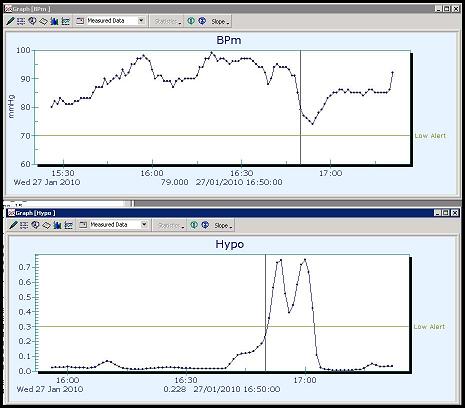
ProjectsFor more detailed information on any of these projects, please use the contact details here. Computational methods of evaluating neurological ICU dataBaseline clinical management of traumatic brain injury (TBI) is an ongoing field of research in medicine. Over the past few decades, drug trials for TBI have proved inconclusive and focus has gradually shifted to issues of patient care and management within the intensive care-unit (ICU) to improve clinical outcome. To assess the various methods of treatment, it is useful to study the data compiled from neurological ICUs and compile an empirical "real-world" guideline of patient care. This project investigates the use of computer science techniques to compile and evaluate neurological ICU data, and compare them against published guidelines, and the statements of adherence by clinicians themselves, with the aim of improving patient outcome. Based in the department of Clinical Physics at the University of Glasgow, and building on data-sets compiled by the Brain-IT consortium, the first goals are to build up a record of treatment instances through explicit annotation or inference of physiological parameter patterns. The project is related to - and uses the infrastructure built in - the Avert-IT project (see also below). Medical motivationTBI treatments and guidelines over the past decades have increasingly focused on the management of critical physiological parameters of a patient, such as intracranial pressure (ICP) and cerebral perfusion pressure (CPP), closely tied to the blood pressure and cerebral blood-flow auto-regulatory mechanisms. Adverse events that can occur, which directly affect these parameters, include secondary insults such as hypotension – where the patient’s blood pressure drops to an abnormally low level – or tachycardia – a dangerously high heart rate. In a typical ICU environment interventions for these events can include: mechanical ventilators, cardiac monitors, external pacemakers, defibrillators, dialysis, intravenous lines/tubes, suction pumps/drains/catheters, and various drugs to induce coma/analgesia/sedation. When studying TBI, neurological ICUs tend to have more focused and specialised equipment. These include: intraventricular drains, ICP monitors, jugular venous bulb oximetry monitors, brain-tissue oximetry, cerebral microdialysis and transcranial Doppler ultrasound. Computer scienceVarious methods are being investigated to study the information patterns that emerge from this environment: knowledge-based ontologies have been compiled and applied to the data-sets to evaluate the physiological output for treatment "signatures". Additionally, various forms of programmable guideline software are being looked at to represent the guideline process in the medical domain (e.g. Asbru, GLIF, Arden syntax). Avert-IT
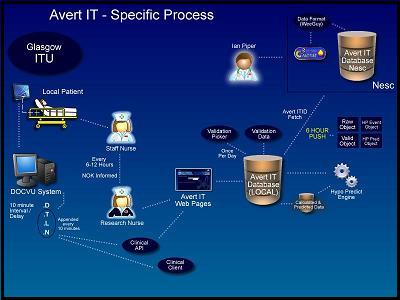
To collect the data, a distributed platform has been created to automatically integrate with bedside monitoring systems within each centre. These include market-leading technologies such as, Philips Docvu, Draeger Medical, Datex-Ohmeda. Every 20 minutes the Avert-IT system interacts with these systems, collates data on patients entered in the study, and using web services, pushes the data to a central repository based at the University of Glasgow.
The clinical trial for this system began at the end of March 2010 and ran for 18 months. ENSAT
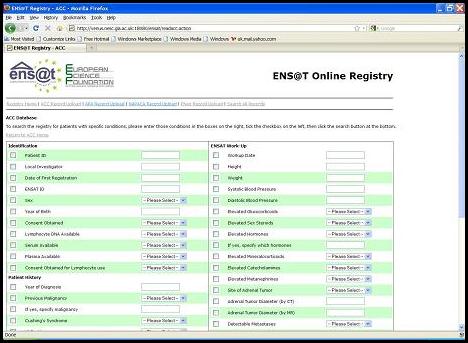
The registry provides interconnected functionality that allows the clinical specialists to collaborate in a way previously not possible, due to administrative issues. A central ID indexes the records across international boundaries, which facilitates the exchange of biomaterial samples. This same index allows the core registry to form a flow of patient information between other related clinical studies (such as PMT - the Prospective Monoamine-producing Tumor study). With ethical approval from all the European nations involved, the registry is an important milestone in the development and communication of rare-disease informatics platforms. |
|||||
|
|
|||||
|
|